 To tackle this issue, we introduce a new algorithm, named SAMOA (Scaffold Constrained Molecular Generation), to perform scaffold-constrained in silico molecular design. Without enforcing such constraints, the probability of generating molecules with the required scaffold is extremely low and hinders the practicality of generative models for de novo drug design. During the optimization of a lead series, it is common to have scaffold constraints imposed on the structure of the molecules designed. One of the major applications of generative models for drug discovery targets the lead-optimization phase. Using predicted descriptors, the fusion model is end-to-end, and requires approximately only 70ms per reaction to predict the selectivity from reaction SMILES strings. For three general types of substitution reactions (aromatic C-H functionalization, aromatic C-X substitution, and other substitution reactions) curated from a commercial database, the fusion model achieves 89:7%, 96:7%, and 97:2% top-1 accuracy in predicting the major outcome, respectively, each using 5000 training reactions. Furthermore, the proposed protocol is demonstrated to be generally applicable to a diverse range of chemical spaces. The proposed platform enhances the inter/extrapolated performance for regio-selectivity predictions and enables learning from small datasets with just hundreds of examples. We construct a reactivity descriptor database based on ab initio calculations of 130k organic molecules, and train a multi-task constrained model to calculate demanded descriptors on-the-fly. We introduce a new method that combines machine-learned reaction representation with selected quantum mechanical descriptorsto predict regio-selectivity in general substitution reactions. Despite the potential of machine learning in predicting chemical reactivity such as selectivity, popular feature engineering and learning methods are either time-consuming or data-hungry. All code is open-source and freely available.Īccurate and rapid evaluation of whether substrates can undergo the desired the transformation iscrucial and challenging for both human knowledge and computer predictions. The new improved generative (DeepFMPO v3D) method is demonstrated with a scaffold-hopping exercise identifying CDK2 bioisosteres. We developed a new python package, ESP-Sim, for the comparison of electrostatic potential and molecular shape, allowing the calculation of state-of-the-art partial charges (e.g., RESP with B3LYP/6-31G**) obtained using the quantum chemistry program Psi4. By performing comparisons of 3D-fragments, we can simulate 3D properties while overcoming the notoriously difficult step of accurately describing bioactive conformations. As before, the generative process outputs optimized molecules similar to the input structures, now with the improved feature of replacing parts of these molecules with fragments of similar 3D-shape and electrostatics. Here, we present a significant extension to our previously described fragment-based reinforcement learning method (DeepFMPO) for the generation of novel molecules with optimal properties. Thus, improved methods for automated compounds design with multiple controlled properties are desired. And I think that your question initially meant to compare the size of #Al^(3+)# NOT #Al^(3-)# (largely unknown).Īnd now to the anions, and cations.and for a given PERIOD, a CATION should be smaller than an anion, inasmuch as we remove an electron, and the remaining electrons should be (and are) held more tightly at closer radii due to the absence of electronic shielding.Multi-parameter optimization, the heart of drug design, is still an open challenge. For selenium, potassium, and bromine, we gots #n=4#.potassium should have the biggest atomic radius, followed by selenium, then bromine.i.e. Than 4th row potassium, selenium, and bromine.because its valence shell is #n=3# rather than #n=4#. 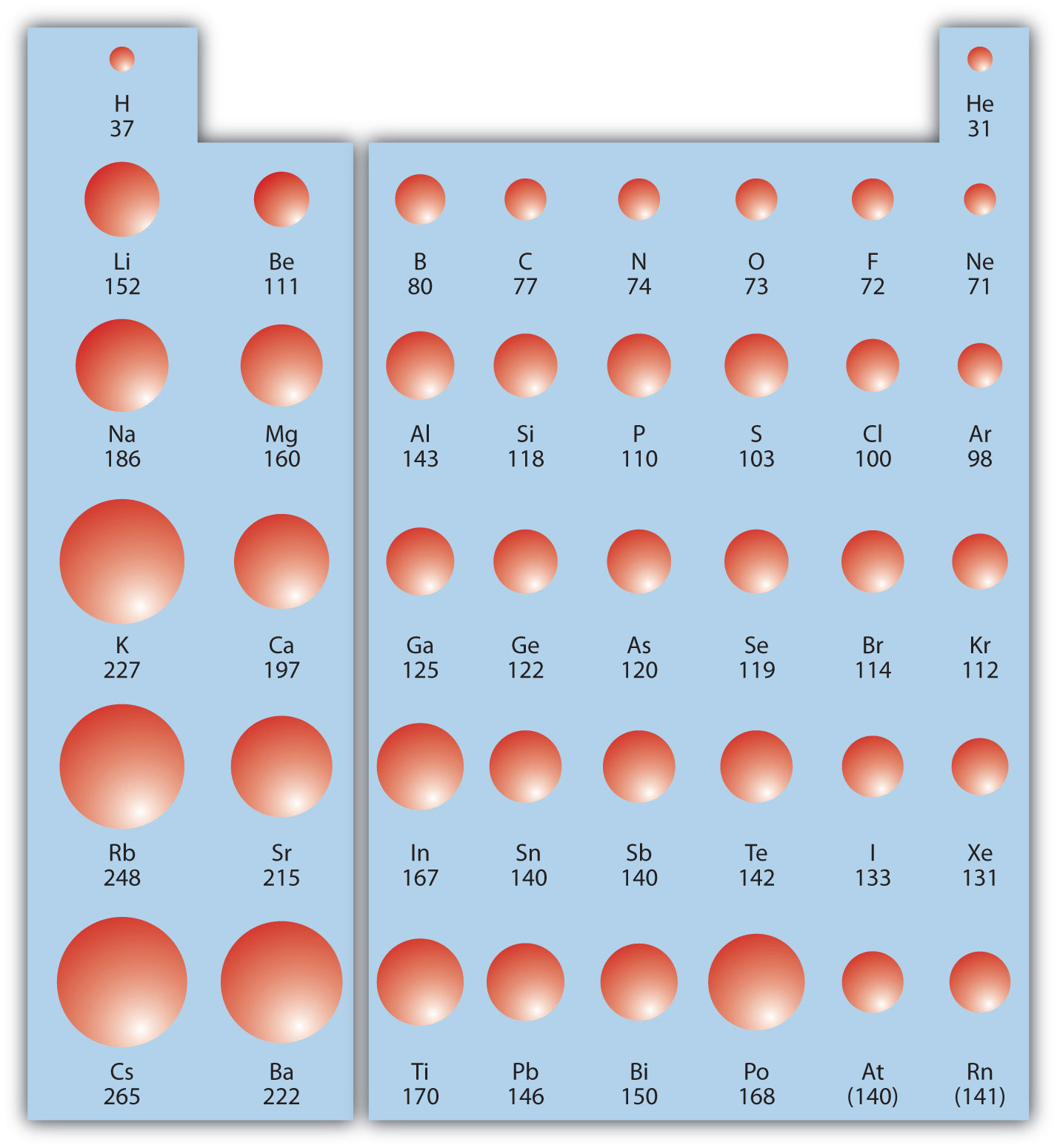
And this is basic and assumed knowledge for 1st year.Īnd so we repair to the Table.and we compare, #Al#, #Se#, #K#, and #Br# ATOMS.Īluminum is a third-row element.and thus it SHOULD be smaller 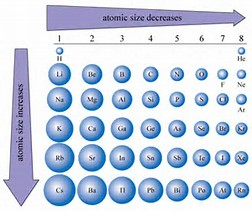
And of course with respect to a row, a Period of the Periodic Table as we face the Table, the size atoms (defined by their valence electronic radius) should DECREASE, and that with respect to a column, a Group of the Periodic Table, atomic radii should INCREASE down the Group. Reasonably, we address the radii of ions and atoms, by the radius of their VALENCE electron. ANIONS should be LARGER than their parent atoms? 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
